 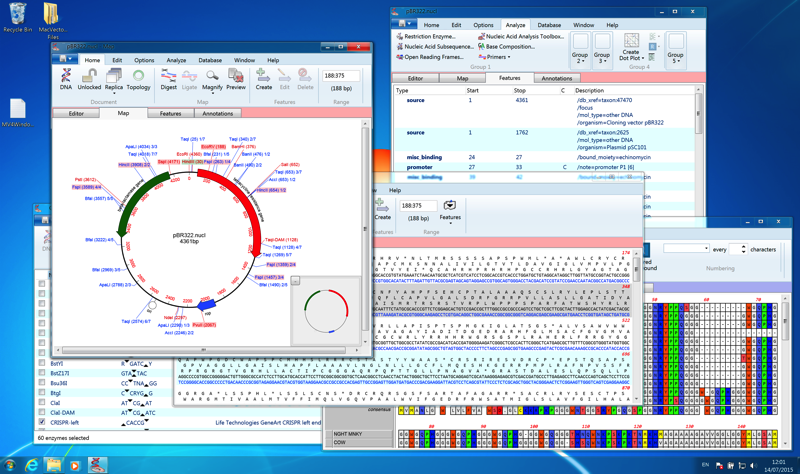
The interactive interface clearly shows which potential primers may have secondary structure problems, alternate binding sites or other characteristics that might impact their use in experiments. MacVector provides a wide variety of useful DNA analysis tools, including base composition analysis, Restriction Enzyme searches, DNA Subsequence searches and "Dot-Plot" comparisons between DNA:DNA and DNA:Protein sequences. A Coding Preference toolbox lets you select a variety of algorithms to graphically scan a DNA sequence for likely protein coding open reading frames. 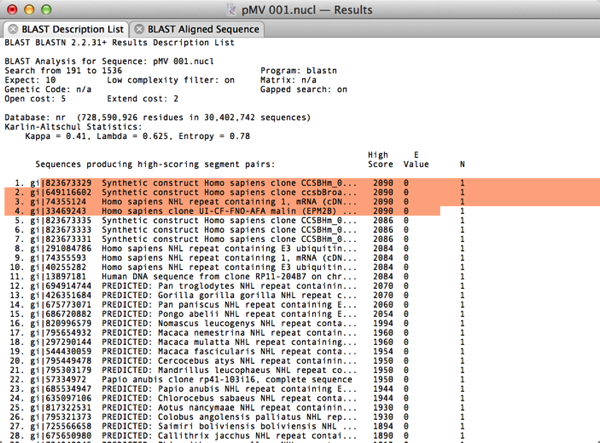
Protein sequences can be reverse translated into DNA, compared using "Dot-Plot" analysis and scanned for Proteolytic cleavage sites and amino acid sequence motifs. A comprehensive Protein Analysis Toolbox provides a wide variety of algorithms for analyzing the composition of proteins and presenting the results in graphical and tabular formats. MacVector has built-in Internet connectivity to the NCBI BLAST and Entrez databases. You can directly search Entrez for DNA or Protein sequences based on features, authors, keywords etc and directly download them into MacVector, complete with all features and annotations. USA 80, 726–730.The built-in BLAST interface lets you submit multiple BLAST jobs using DNA or Protein sequences and then download any matching sequences by selecting them from a hit list. (1983) Rapid similarity searches of nucleic acid and protein databanks. (1990) Rapid and sensitive sequence comparison with FASTP and FASTA. (1985) Rapid and sensitive protein similarity searches. (1982) A high speed, high capacity homology matrix zooming through SV40 and polyoma. (1994) Issues in searching molecular sequence databases. (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, positions-specific gap penalties and weight matrix choice. (1978) Analysis of the accuracy and implications of simple methods for predicting the secondary structure of globular proteins. 
(1974) Conformational parameters for amino acids in helical, beta-sheet, and random coil regions calculated from proteins. (1989) A computer program for choosing optimal oligonucleotides for filter hybridization, sequencing and in vitro amplification of DNA. (1984) The codon preference plot graphic analysis of protein coding sequences and prediction of gene expression. Gribskov, M., Devereux, J., and Burgess, R. (1982) Recognition of protein coding regions in DNA sequences. Oxford Molecular, Oxford, England.įickett, J. Oxford Molecular Group (1998) MacVector 6.5 User Guide. This process is experimental and the keywords may be updated as the learning algorithm improves. These keywords were added by machine and not by the authors. MacVector also comes with a module for contig assembly, called AssemblyLIGN. At the time of writing, the version of MacVector available was 6.5. It provides all of the most commonly used nucleic acid and protein analysis tools and also provides access via the Internet to the public Entrez databases at the National Center for Biotechnology Information (NCBI). It is an integrated comprehensive sequence analysis program that runs on the Macintosh. MacVector™, from Oxford Molecular Group, Campbell CA, is one such computer package. It is evident by now that efficiently performing the above operations on thousands or millions of base pairs by hand is so difficult as to be impossible, and computer programs that do sequence analysis are becoming more and more ubiquitous in laboratories practicing molecular biology. Whether a researcher is working on a genome project, or is cloning and characterizing a gene of interest, the ability to manipulate, analyze, and annotate sequence data is becoming increasingly important. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
